tags:
- biology
- small-moelcule
- single-cell-genes
- ibm
- mammal
- pytorch
- transformers
library_name: biomed
license: apache-2.0
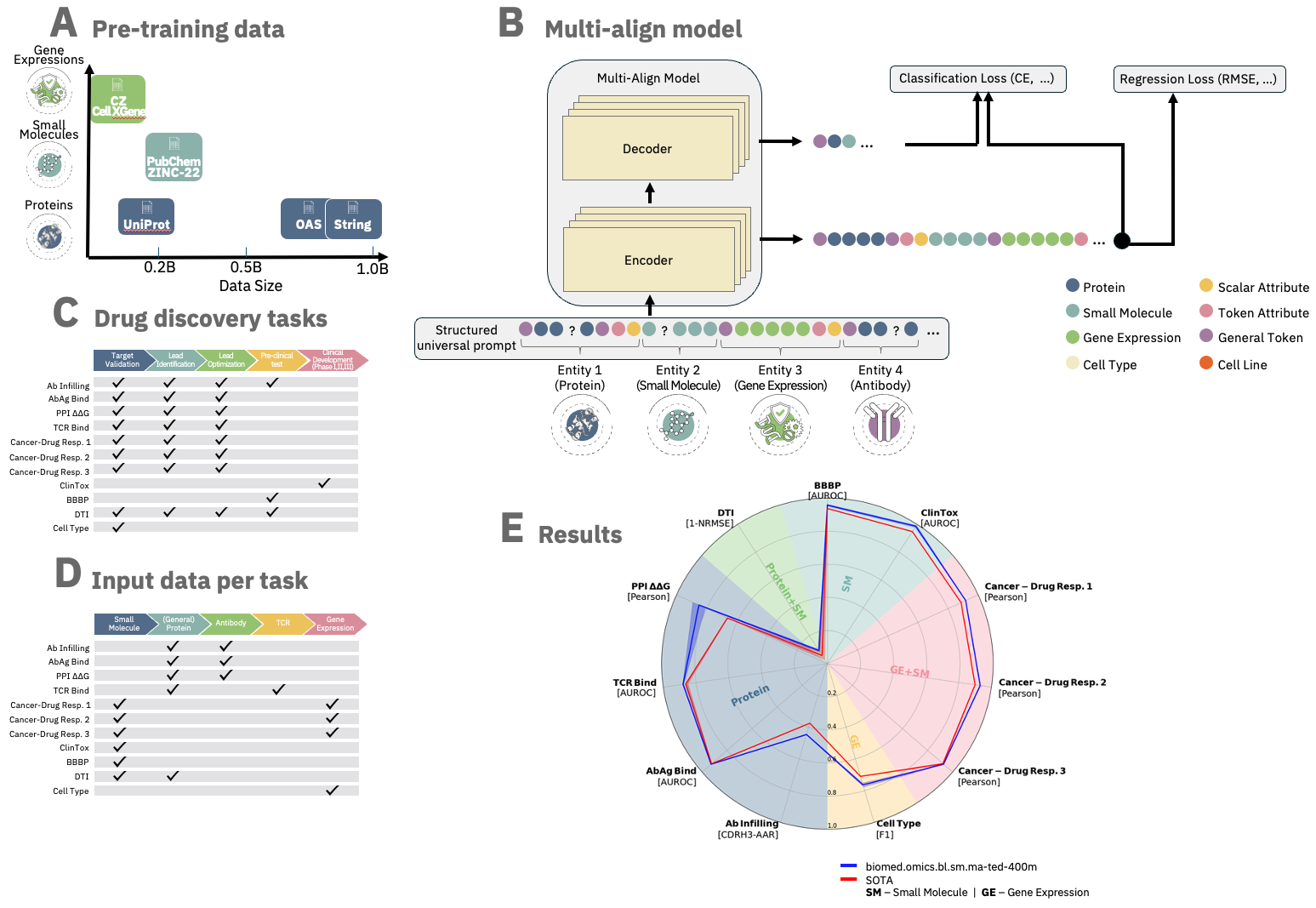
The ibm/biomed.omics.bl.sm.ma-ted-400m model is a biomedical foundation model trained on over 2 billion biological samples across multiple modalities, including proteins, small molecules, and single-cell gene data.
Designed for robust performance, it achieves state-of-the-art results over a variety of tasks across the entire drug discovery pipeline and the diverse biomedical domains.
Based on the Molecular Aligned Multi-Modal Architecture and Language (MAMMAL), a flexible, multi-domain architecture with an adaptable task prompt syntax.
The syntax allows for dynamic combinations of tokens and scalars, enabling classification, regression, and generation tasks either within a single domain or with cross-domain entities.
Model Summary
- Developers: IBM Research
- GitHub Repository: https://github.com/BiomedSciAI/biomed-multi-alignment
- Paper: TBD
- Release Date: Oct 28th, 2024
- License: Apache 2.0.
Usage
Using ibm/biomed.omics.bl.sm.ma-ted-400m requires installing https://github.com/BiomedSciAI/biomed-multi-alignment
pip install git+https://github.com/BiomedSciAI/biomed-multi-alignment.git
A simple example for a task already supported by ibm/biomed.omics.bl.sm.ma-ted-400m:
import torch
from fuse.data.tokenizers.modular_tokenizer.op import ModularTokenizerOp
from mammal.model import Mammal
from mammal.keys import *
# Load Model
model = Mammal.from_pretrained("ibm/biomed.omics.bl.sm.ma-ted-400m")
# Load Tokenizer
tokenizer_op = ModularTokenizerOp.from_pretrained("ibm/biomed.omics.bl.sm.ma-ted-400m")
# Prepare Input Prompt
protein_calmodulin = "MADQLTEEQIAEFKEAFSLFDKDGDGTITTKELGTVMRSLGQNPTEAELQDMISELDQDGFIDKEDLHDGDGKISFEEFLNLVNKEMTADVDGDGQVNYEEFVTMMTSK"
protein_calcineurin = "MSSKLLLAGLDIERVLAEKNFYKEWDTWIIEAMNVGDEEVDRIKEFKEDEIFEEAKTLGTAEMQEYKKQKLEEAIEGAFDIFDKDGNGYISAAELRHVMTNLGEKLTDEEVDEMIRQMWDQNGDWDRIKELKFGEIKKLSAKDTRGTIFIKVFENLGTGVDSEYEDVSKYMLKHQ"
# Create and load sample
sample_dict = dict()
# Formatting prompt to match pre-training syntax
sample_dict[ENCODER_INPUTS_STR] = f"<@TOKENIZER-TYPE=AA><BINDING_AFFINITY_CLASS><SENTINEL_ID_0><MOLECULAR_ENTITY><MOLECULAR_ENTITY_GENERAL_PROTEIN><SEQUENCE_NATURAL_START>{protein_calmodulin}<SEQUENCE_NATURAL_END><MOLECULAR_ENTITY><MOLECULAR_ENTITY_GENERAL_PROTEIN><SEQUENCE_NATURAL_START>{protein_calcineurin}<SEQUENCE_NATURAL_END><EOS>"
# Tokenize
tokenizer_op(
sample_dict=sample_dict,
key_in=ENCODER_INPUTS_STR,
key_out_tokens_ids=ENCODER_INPUTS_TOKENS,
key_out_attention_mask=ENCODER_INPUTS_ATTENTION_MASK,
)
sample_dict[ENCODER_INPUTS_TOKENS] = torch.tensor(sample_dict[ENCODER_INPUTS_TOKENS])
sample_dict[ENCODER_INPUTS_ATTENTION_MASK] = torch.tensor(sample_dict[ENCODER_INPUTS_ATTENTION_MASK])
# Generate Prediction
batch_dict = model.generate(
[sample_dict],
output_scores=True,
return_dict_in_generate=True,
max_new_tokens=5,
)
# Get output
generated_output = tokenizer_op._tokenizer.decode(batch_dict[CLS_PRED][0])
print(f"{generated_output=}")
For more advanced usage, see our detailed example at:
Citation
If you found our work useful, please consider to give a star to the repo and cite our paper:
@article{TBD,
title={TBD},
author={IBM Research Team},
jounal={arXiv preprint arXiv:TBD},
year={2024}
}